Difference between revisions of "Main Page"
(1.7.4) |
(PyMOL v1.8.2) |
||
| (9 intermediate revisions by 4 users not shown) | |||
| Line 17: | Line 17: | ||
| style="font-size: 1.1em; color #61021F; padding: 0.5em 1em 0.5em 3em;"|'''[[Gallery]]''' | '''[[Covers]]''' | | style="font-size: 1.1em; color #61021F; padding: 0.5em 1em 0.5em 3em;"|'''[[Gallery]]''' | '''[[Covers]]''' | ||
||'''[[CheatSheet|PyMOL Cheat Sheet]]''' (''[[Media:PymolRef.pdf|PDF]]'') | ||'''[[CheatSheet|PyMOL Cheat Sheet]]''' (''[[Media:PymolRef.pdf|PDF]]'') | ||
| − | ||'''[[ | + | ||'''[[PyMOL_mailing_list|Getting Help]]''' |
|} | |} | ||
| Line 26: | Line 26: | ||
|- | |- | ||
! Official Release | ! Official Release | ||
| − | | [http://pymol.org PyMOL | + | | [http://pymol.org PyMOL v1.8.2 has been released] on April 20, 2016. |
| + | |- | ||
| + | ! New Script | ||
| + | | [[dssr_block]] is a wrapper for DSSR (3dna) and creates block-shaped nucleic acid cartoons | ||
|- | |- | ||
! New Plugin | ! New Plugin | ||
| − | | [[ | + | | [[Lisica|LiSiCA]] is a new plugin for 2D and 3D ligand based virtual screening using a fast maximum clique algorithm. |
| + | |- | ||
| + | ! Official Release | ||
| + | | [http://pymol.org PyMOL v1.8.0 has been released] on Nov 18, 2015. | ||
|- | |- | ||
! PyMOL Open-Source Fellowship | ! PyMOL Open-Source Fellowship | ||
| Line 35: | Line 41: | ||
|- | |- | ||
! Official Release | ! Official Release | ||
| − | | [http://pymol.org PyMOL, AxPyMOL, and JyMOL v1.7. | + | | [http://pymol.org PyMOL, AxPyMOL, and JyMOL v1.7.6 have all been released] on May 4, 2015. |
|- | |- | ||
| − | ! | + | ! New Plugin |
| − | | [ | + | | [[PyANM|PyANM]] is a new plugin for easier Anisotropic Network Model (ANM) building and visualising in PyMOL. |
|- | |- | ||
! New Plugin | ! New Plugin | ||
| Line 48: | Line 54: | ||
! 3D using Geforce | ! 3D using Geforce | ||
| PyMOL can now be [http://forums.geforce.com/default/topic/571604/3d-vision/3d-vision-working-with-qbs-in-opengl-software-using-geforce/2/ visualized in 3D using Nvidia GeForce video cards] (series 400+) with 120Hz monitors and Nvidia 3D Vision, this was previously only possible with Quadro video cards. | | PyMOL can now be [http://forums.geforce.com/default/topic/571604/3d-vision/3d-vision-working-with-qbs-in-opengl-software-using-geforce/2/ visualized in 3D using Nvidia GeForce video cards] (series 400+) with 120Hz monitors and Nvidia 3D Vision, this was previously only possible with Quadro video cards. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
|- | |- | ||
! Older News | ! Older News | ||
| Line 69: | Line 63: | ||
|- | |- | ||
|<div class="didyouknow" > | |<div class="didyouknow" > | ||
| − | < | + | <DynamicPageList> |
| − | + | randomcount=1 | |
category=Commands|Plugins|Script_Library|Settings | category=Commands|Plugins|Script_Library|Settings | ||
includepage=* | includepage=* | ||
includemaxlength=1050 | includemaxlength=1050 | ||
escapelinks=false | escapelinks=false | ||
| + | allowcachedresults=false | ||
resultsheader=__NOTOC__ __NOEDITSECTION__ | resultsheader=__NOTOC__ __NOEDITSECTION__ | ||
| − | |||
| − | |||
| − | |||
listseparators=,<h3>[[%PAGE%]]</h3>,,\n | listseparators=,<h3>[[%PAGE%]]</h3>,,\n | ||
| − | </ | + | </DynamicPageList> |
</div> | </div> | ||
<div style="clear: both;"></div> | <div style="clear: both;"></div> | ||
| Line 86: | Line 78: | ||
| | | | ||
|style="vertical-align: top; width: 18%"| | |style="vertical-align: top; width: 18%"| | ||
| − | < | + | <DynamicPageList> |
imagecontainer=Covers | imagecontainer=Covers | ||
randomcount=1 | randomcount=1 | ||
| Line 93: | Line 85: | ||
listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]],\n | listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]],\n | ||
ordermethod=none | ordermethod=none | ||
| − | </ | + | allowcachedresults=false |
| + | </DynamicPageList> | ||
|} | |} | ||
Revision as of 10:57, 20 April 2016
| The community-run support site for the PyMOL molecular viewer. |
| New accounts: email jason (dot) vertrees (@) gmail dot com |
| Tutorials | Table of Contents | Commands |
| Script Library | Plugins | FAQ |
| Gallery | Covers | PyMOL Cheat Sheet (PDF) | Getting Help |
|
|
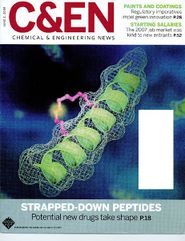 A Random PyMOL-generated Cover. See Covers.
| ||||||||||||||||||||||||||||||||||