AAindex
Jump to navigation
Jump to search
| Type | Python Script |
|---|---|
| Download | scripts/aaindex.py |
| Author(s) | Thomas Holder |
| License | BSD |
| This code has been put under version control in the project Pymol-script-repo | |
|
Included in psico | |
| Module | psico.aaindex |
|---|---|
Introduction
AAindex is a database of numerical indices representing various physicochemical and biochemical properties of amino acids and pairs of amino acids. See http://www.genome.jp/aaindex/
This script is a python parser for the AAindex flat files which will be downloaded from ftp://ftp.genome.jp/pub/db/community/aaindex/
The script provides two PyMOL commands (but can also be used without PyMOL).
- aaindex2b: Loads numerical indices from aaindex1 as b-factors into your structure
- pmf: Potential of Mean Force (aaindex3)
Python Example
Consider the script is called aaindex.py, it is placed somewhere in your PYTHONPATH and the aaindex flatfiles are found in the current directory.
import aaindex
aaindex.init(path='.')
aaindex.grep('volume')
x = aaindex.get('KRIW790103')
print(x)
print(x.get('A'))
aaindex.init(index='2')
aaindex.grep('blosum')
x = aaindex.get('HENS920102')
print(x.get('A', 'K'))
PyMOL Example
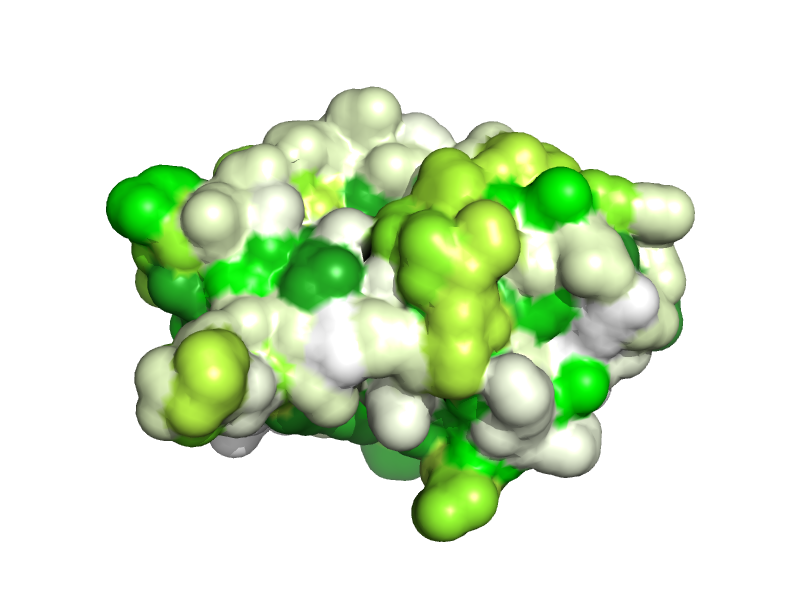
Solvent-accessible surface coloring by Hydropathy index (Kyte-Doolittle, 1982)
run aaindex.py
aaindex2b KYTJ820101
spectrum b, white yellow forest
set surface_solvent
show surface
See Also
- Protscale from rTools does a similar job in coloring by amino acid properties