Difference between revisions of "Color by conservation"
Jump to navigation
Jump to search
m (d'oh) |
|||
| Line 1: | Line 1: | ||
| + | {{Infobox script-repo | ||
| + | |type = script | ||
| + | |filename = color_by_conservation.py | ||
| + | |author = [[Users:Inchoate|Jason Vertrees]] | ||
| + | |license = Free | ||
| + | }} | ||
| + | |||
= Overview = | = Overview = | ||
This script reads an alignment object and colors the protein objects in the alignment by the sequence conservation found in the alignment. | This script reads an alignment object and colors the protein objects in the alignment by the sequence conservation found in the alignment. | ||
| Line 5: | Line 12: | ||
= Example Usage = | = Example Usage = | ||
| − | < | + | <syntaxhighlight lang="python"> |
| − | # | + | reinitialize |
| − | + | import color_by_conservation | |
| − | + | ||
| − | + | # get some kinases | |
| + | fetch 1opk 3dtc 3p86 2eva 3efw, async=0 | ||
| + | |||
# turn on the sequence viewer | # turn on the sequence viewer | ||
| − | |||
set seq_view | set seq_view | ||
| − | + | # align them into the "algn" object | |
| − | + | for x in cmd.get_names(): cmd.align(x, "3efw and c. A", object="algn") | |
| − | + | ||
| − | |||
| − | # align them into the " | ||
| − | |||
| − | for x in cmd.get_names(): cmd.align(x, "3efw and c. A", object=" | ||
| − | |||
# color | # color | ||
| − | + | color_by_conservation aln=algn, as_putty=1 | |
| − | color_by_conservation | + | </syntaxhighlight> |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | </ | ||
[[Category:Script_Library]] | [[Category:Script_Library]] | ||
[[Category:Structural_Biology_Scripts]] | [[Category:Structural_Biology_Scripts]] | ||
Revision as of 11:27, 14 December 2011
| Type | Python Script |
|---|---|
| Download | color_by_conservation.py |
| Author(s) | Jason Vertrees |
| License | Free |
| This code has been put under version control in the project Pymol-script-repo | |
Overview
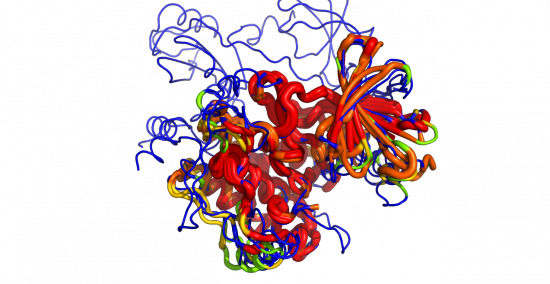
This script reads an alignment object and colors the protein objects in the alignment by the sequence conservation found in the alignment.
Example Usage
reinitialize
import color_by_conservation
# get some kinases
fetch 1opk 3dtc 3p86 2eva 3efw, async=0
# turn on the sequence viewer
set seq_view
# align them into the "algn" object
for x in cmd.get_names(): cmd.align(x, "3efw and c. A", object="algn")
# color
color_by_conservation aln=algn, as_putty=1