Cealign
Appearance
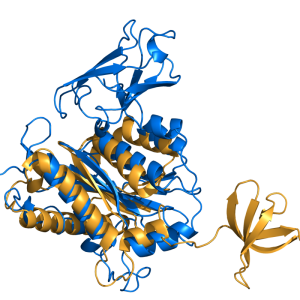
cealign aligns two proteins using the CE algorithm. It is very robust for proteins with little to no sequence similarity (twilight zone). For proteins with decent structural similarity, the super command is preferred and with decent sequence similarity, the align command is preferred, because these commands are much faster than cealign.
This command is new in PyMOL 1.3, see the cealign plugin for manual installation.
Usage
cealign target, mobile [, target_state [, mobile_state
[, quiet [, guide [, d0 [, d1 [, window [, gap_max
[, transform [, object ]]]]]]]]]]
Arguments
Note: The mobile and target arguments are swapped with respect to the align and super commands.
- target = string: atom selection of target object (CE uses only CA atoms)
- mobile = string: atom selection of mobile object (CE uses only CA atoms)
- target_state = int: object state of target selection {default: 1}
- mobile_state = int: object state of mobile selection {default: 1}
- quiet = 0/1: suppress output {default: 0 in command mode, 1 in API}
- guide = 0/1: only use "guide" atoms (CA, C4') {default: 1}
- d0, d1, window, gap_max: CE algorithm parameters
- transform = 0/1: do superposition {default: 1}
- object = string: name of alignment object to create {default: (no alignment object)}
Example
fetch 1c0mB 1bco, async=0
as ribbon
cealign 1bco, 1c0mB, object=aln