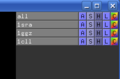
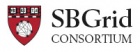command creates or updates a "group" object. The grouped objects are collected underneath a
sign in the object tree (see images) in the Pymol Internal Gui.
# put all of objState into the group "ensemble".
group ensemble, objState*
group name, members, action
group efHand, 1cll 1ggz 1sra
# allow addition and removal from the group
# If a group is open, objects can be added to or removed from
# it by right-click+drag from the control panel
group efHand, open
# disallow addition/removal from the group
group efHand, close
# names with dots are treated special
set group_auto_mode, 2
# load the example protein
load $TUT/1hpv.pdb, 1hpv.other
# create the new entry called ".protein" in group 1hpv
extract 1hpv.protein, 1hpv.other and polymer
# create ".ligand in the 1hpv group
extract 1hpv.ligand, 1hpv.other and organic
# supports wildcards
show sticks, *.ligand
hide lines, *.protein
show surface, *.protein within 6 of *.ligand
show lines, byres *.protein within 4 of *.ligand
set two_sided_lighting
set transparency, 0.5
set surface_color, white
# Also, to lexicographically sort the names in the control panel:
order *, yes
Group objects can usually be used as arguments to commands. It can be processed as a group or as a selection, in which case all the atoms from all objects in the group will be used.