Difference between revisions of "Bnitools"
Jump to navigation
Jump to search
(Created page with "{{Infobox script-repo |type = plugin |filename = plugin/bnitools.py |author = Georg Steinkellner |license = BSD }}") |
|||
| (11 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
{{Infobox script-repo | {{Infobox script-repo | ||
|type = plugin | |type = plugin | ||
| − | |filename = | + | |filename = plugins/bnitools.py |
| − | |author = [[User: | + | |author = [[User:Steinkeg|Georg Steinkellner]] |
|license = BSD | |license = BSD | ||
}} | }} | ||
| + | |||
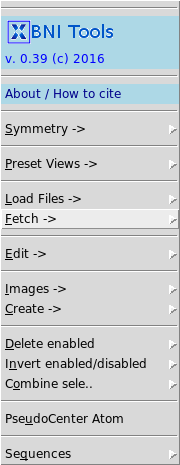
| + | [[File:bnitools.png||thumb|right|Screenshot of the BNI Tools plugin menu]] | ||
| + | |||
| + | '''BNI Tools''' is a plug in for PyMOL which adds additional functionalities and presets to the PyMOL GUI. | ||
| + | |||
| + | == Installation == | ||
| + | |||
| + | Install it using the 'Install Plugin' menu within PyMOL:<br> | ||
| + | '''PyMOL > Plugin > Plugin Manager'''<br> | ||
| + | |||
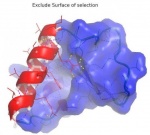
| + | == Example images created using BNI Tools == | ||
| + | [[File:bni_box.jpg|150px|||Create a box around a selection]] | ||
| + | [[File:bni_delphimap.jpg|150px|||Load and show DelPhi or other maps]] | ||
| + | [[File:bni_box2.jpg|150px|||Create a box]] | ||
| + | [[File:Bni_excludesurf.jpg|150px|||Exclude parts of surface]] | ||
| + | [[File:Bni_plane.jpg|150px|||Create Plane]] | ||
| + | [[File:Bni_track.jpg|150px|||Create a main chain track]] | ||
| + | == BNI Tools dropdown menu (Plugin > BNI Tools > ...) == | ||
| + | |||
| + | [+]Load Files-> | ||
| + | [-]modeller files | ||
| + | * load multiple modeller files and sort by | ||
| + | * objective function energy | ||
| + | * and add energy to object title | ||
| + | [-]autodock files | ||
| + | * load autodock .dlg or .dlg.pdb file into | ||
| + | * different states with energy in object | ||
| + | * title | ||
| + | [-]amber minimization | ||
| + | * load amber minimization file with energy in | ||
| + | * title if the .info file is present in same | ||
| + | * directory and has the same name as the pdb file. | ||
| + | [-]delphi phi,dx map | ||
| + | * load delphi map and corresponding pdb file | ||
| + | * simultaneously and show surface colored by | ||
| + | * PHI or DX map. (show the surface to see the | ||
| + | * effect) | ||
| + | [-]casox map | ||
| + | * load casox map (cavity calculation) and | ||
| + | * show ligsite accessibility | ||
| + | * value maps. (7 closed cavity to 1 open) | ||
| + | [-]multiple files into states | ||
| + | * load multiple pdb files (e.g. MD simulation | ||
| + | * snapshots) into one state. (object is named | ||
| + | * by first object loaded) | ||
| + | [+]Fetch-> | ||
| + | [-]Density View (EDS) | ||
| + | * load density and pdb file from EDS | ||
| + | * (Electron Density Server) | ||
| + | * if available, and show density with density | ||
| + | * wizard | ||
| + | [-]RCSB Biol. Assembly | ||
| + | * load biological assembly from RCSB | ||
| + | * protein database | ||
| + | [-]2FoFc map(s) | ||
| + | * load (multiple) 2FoFc maps | ||
| + | * from EDS density server | ||
| + | * if available | ||
| + | [-]FoFc maps(s) | ||
| + | * load (multiple) FoFc maps | ||
| + | * from EDS density server | ||
| + | * if available | ||
| + | [+]Edit-> | ||
| + | [-]HIS --> HID,HIE,HIP | ||
| + | * change histidine residues to HID,HIE,HIP | ||
| + | * depending on hydrogens on histidine | ||
| + | [-]HID,HIE,HIP --> HIS | ||
| + | * change altered histidine residues | ||
| + | * back to HIS | ||
| + | [-]Poly-Alanine Chain | ||
| + | * create a poly alanine chain (GLY and ALA) | ||
| + | * for molecular replacement | ||
| + | [-]MSE --> MET | ||
| + | * change selenomethionine to methionine | ||
| + | [-]del alternates | ||
| + | * delete alternates in selection | ||
| + | [-]Unbond- > | ||
| + | * unbond atoms in selection | ||
| + | [+]Images-> | ||
| + | * create ray traced images depending on | ||
| + | * x size and resolution (dpi) | ||
| + | [+]Create-> | ||
| + | * create compiled graphics objects (CGO) | ||
| + | * these objects can be altered in color or | ||
| + | * transparency, and they can be dragged | ||
| + | * and rotated in space by the | ||
| + | * "action->drag" command and using | ||
| + | * "shift" and mouse buttons. | ||
| + | [-]Plane | ||
| + | * create a plane (with certain cushion) | ||
| + | * using a selection of three atoms | ||
| + | [-]Box | ||
| + | * create a box around selection | ||
| + | * the whole box can be altered as group | ||
| + | * or by side planes separately | ||
| + | [-]Triangle | ||
| + | * create a triangle using a selection of three | ||
| + | * atoms | ||
| + | [+]pseudo center atom | ||
| + | * create pseudo atom at the center of atoms | ||
| + | * in selection | ||
| + | * move atoms with editing->"shift" | ||
| + | * and middle mouse button | ||
| + | |||
| + | == BNI tools integrated in PyMOL sidebar == | ||
| + | |||
| + | [+]Action on "all" | ||
| + | [-]delete enabled | ||
| + | * delete all enabled objects or | ||
| + | * selections | ||
| + | [-]invert enabled/disabled | ||
| + | * disable currently enabled objects | ||
| + | [-]combine selections | ||
| + | * combine all enabled or disabled | ||
| + | * selections to selection (sele) | ||
| + | [+]action | ||
| + | [-]sequence | ||
| + | * show sequence in different formats | ||
| + | * of selection | ||
| + | * copy and paste to text file for later use | ||
| + | [.]fasta | ||
| + | * show sequence of selection | ||
| + | * in fasta format | ||
| + | [.]pir | ||
| + | * show sequence of selection | ||
| + | * in pir format | ||
| + | [.]modeller | ||
| + | * show sequence of selection | ||
| + | * in modeller pir format | ||
| + | [.]list | ||
| + | * create residue or atom lists | ||
| + | * of selection | ||
| + | [+]preset | ||
| + | [-]track main chain | ||
| + | * create a new object which tracks the | ||
| + | * main chain atoms and shows main chain | ||
| + | * and side chain polar contacts | ||
| + | [-]symmetry surface | ||
| + | * create a symmetry view of the selected | ||
| + | * atoms showing the contact surface as well | ||
| + | * as a selection entry of the atoms in | ||
| + | * contact with symmetry mates | ||
| + | * (only includes atoms of the initial selection). | ||
| + | [-]hydrophobic residues | ||
| + | * show hydrophobic residues | ||
| + | * depending on hydrophobic residue scales | ||
| + | * by KandD (Kyte & Doolittle | ||
| + | * J Mol Biol 157:105, 1982) | ||
| + | * Rose (Rose et al. | ||
| + | * Science, 229, 834-838,1985) | ||
| + | * GES (Engelman Engelman et al. | ||
| + | * Annu Rev Biophys Biophys Chem, | ||
| + | * 15, 321-353(1986) | ||
| + | * (no window selection; just raw categories | ||
| + | * are colored by: blue-hydrophile | ||
| + | * green-neutral | ||
| + | * red-hydrophobe ) | ||
| + | [-]surface inspection | ||
| + | * create selections for surface | ||
| + | * inspections | ||
| + | # BNI tools additional settings in PyMOL sidebar | ||
| + | [+]color | ||
| + | [-]by ss | ||
| + | * color helix,sheet and loop separately | ||
| + | [.]Helix | ||
| + | [.]Sheet | ||
| + | [.]Loop | ||
| + | # this section is replaced by "[-] by_rep" in PyMOL versions >1.6 | ||
| + | [-]surface | ||
| + | * color surface separately from atoms | ||
| + | [.]by atom | ||
| + | * set surface color to standard | ||
| + | [.]by map | ||
| + | * if a map or ramp is loaded | ||
| + | * color surface by ramp/map | ||
| + | [.]my color | ||
| + | * color surface by own defined color | ||
| + | [-]mesh | ||
| + | * color mesh separately from atoms | ||
| + | [.]by atom | ||
| + | * set mesh color to standard | ||
| + | [.]by map | ||
| + | * if a map or ramp is loaded | ||
| + | * color mesh by ramp/map | ||
| + | [.]my color | ||
| + | * color mesh by own defined color | ||
| + | [-]label | ||
| + | * color labels separately | ||
| + | [.]by atom | ||
| + | * set label color by atom | ||
| + | [.]my color | ||
| + | * color labels by own defined color | ||
| + | [-]stick | ||
| + | * color sticks separately | ||
| + | [.]standard | ||
| + | * set stick color to standard | ||
| + | [.]my color | ||
| + | * color sticks by own defined color | ||
| + | [-]my colors | ||
| + | * use/append own defined colors | ||
| + | * own colors can be defined by | ||
| + | * Setting->Colors..->New | ||
| + | * to keep color settings for | ||
| + | * other pymol sessions | ||
| + | * you have to set the colors | ||
| + | * in .pymolrc or similar | ||
| + | * pymol setting file | ||
| + | * like | ||
| + | * set_color mycolor,[ 1.00, 1.00, 1.00] | ||
| + | [+]show | ||
| + | [-]surface flag | ||
| + | * set surface flag of atoms to show hide | ||
| + | * or ignore fro surface calculation | ||
| + | [-]transparency | ||
| + | * set different transparency types | ||
| + | * on selections or atoms | ||
| + | |||
| + | [[Category:Plugins]] | ||
| + | [[Category:Pymol-script-repo]] | ||
Latest revision as of 10:45, 12 May 2016
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/bnitools.py |
| Author(s) | Georg Steinkellner |
| License | BSD |
| This code has been put under version control in the project Pymol-script-repo | |
BNI Tools is a plug in for PyMOL which adds additional functionalities and presets to the PyMOL GUI.
Installation
Install it using the 'Install Plugin' menu within PyMOL:
PyMOL > Plugin > Plugin Manager
Example images created using BNI Tools
[+]Load Files->
[-]modeller files
* load multiple modeller files and sort by
* objective function energy
* and add energy to object title
[-]autodock files
* load autodock .dlg or .dlg.pdb file into
* different states with energy in object
* title
[-]amber minimization
* load amber minimization file with energy in
* title if the .info file is present in same
* directory and has the same name as the pdb file.
[-]delphi phi,dx map
* load delphi map and corresponding pdb file
* simultaneously and show surface colored by
* PHI or DX map. (show the surface to see the
* effect)
[-]casox map
* load casox map (cavity calculation) and
* show ligsite accessibility
* value maps. (7 closed cavity to 1 open)
[-]multiple files into states
* load multiple pdb files (e.g. MD simulation
* snapshots) into one state. (object is named
* by first object loaded)
[+]Fetch->
[-]Density View (EDS)
* load density and pdb file from EDS
* (Electron Density Server)
* if available, and show density with density
* wizard
[-]RCSB Biol. Assembly
* load biological assembly from RCSB
* protein database
[-]2FoFc map(s)
* load (multiple) 2FoFc maps
* from EDS density server
* if available
[-]FoFc maps(s)
* load (multiple) FoFc maps
* from EDS density server
* if available
[+]Edit->
[-]HIS --> HID,HIE,HIP
* change histidine residues to HID,HIE,HIP
* depending on hydrogens on histidine
[-]HID,HIE,HIP --> HIS
* change altered histidine residues
* back to HIS
[-]Poly-Alanine Chain
* create a poly alanine chain (GLY and ALA)
* for molecular replacement
[-]MSE --> MET
* change selenomethionine to methionine
[-]del alternates
* delete alternates in selection
[-]Unbond- >
* unbond atoms in selection
[+]Images->
* create ray traced images depending on
* x size and resolution (dpi)
[+]Create->
* create compiled graphics objects (CGO)
* these objects can be altered in color or
* transparency, and they can be dragged
* and rotated in space by the
* "action->drag" command and using
* "shift" and mouse buttons.
[-]Plane
* create a plane (with certain cushion)
* using a selection of three atoms
[-]Box
* create a box around selection
* the whole box can be altered as group
* or by side planes separately
[-]Triangle
* create a triangle using a selection of three
* atoms
[+]pseudo center atom
* create pseudo atom at the center of atoms
* in selection
* move atoms with editing->"shift"
* and middle mouse button
BNI tools integrated in PyMOL sidebar
[+]Action on "all"
[-]delete enabled
* delete all enabled objects or
* selections
[-]invert enabled/disabled
* disable currently enabled objects
[-]combine selections
* combine all enabled or disabled
* selections to selection (sele)
[+]action
[-]sequence
* show sequence in different formats
* of selection
* copy and paste to text file for later use
[.]fasta
* show sequence of selection
* in fasta format
[.]pir
* show sequence of selection
* in pir format
[.]modeller
* show sequence of selection
* in modeller pir format
[.]list
* create residue or atom lists
* of selection
[+]preset
[-]track main chain
* create a new object which tracks the
* main chain atoms and shows main chain
* and side chain polar contacts
[-]symmetry surface
* create a symmetry view of the selected
* atoms showing the contact surface as well
* as a selection entry of the atoms in
* contact with symmetry mates
* (only includes atoms of the initial selection).
[-]hydrophobic residues
* show hydrophobic residues
* depending on hydrophobic residue scales
* by KandD (Kyte & Doolittle
* J Mol Biol 157:105, 1982)
* Rose (Rose et al.
* Science, 229, 834-838,1985)
* GES (Engelman Engelman et al.
* Annu Rev Biophys Biophys Chem,
* 15, 321-353(1986)
* (no window selection; just raw categories
* are colored by: blue-hydrophile
* green-neutral
* red-hydrophobe )
[-]surface inspection
* create selections for surface
* inspections
# BNI tools additional settings in PyMOL sidebar
[+]color
[-]by ss
* color helix,sheet and loop separately
[.]Helix
[.]Sheet
[.]Loop
# this section is replaced by "[-] by_rep" in PyMOL versions >1.6
[-]surface
* color surface separately from atoms
[.]by atom
* set surface color to standard
[.]by map
* if a map or ramp is loaded
* color surface by ramp/map
[.]my color
* color surface by own defined color
[-]mesh
* color mesh separately from atoms
[.]by atom
* set mesh color to standard
[.]by map
* if a map or ramp is loaded
* color mesh by ramp/map
[.]my color
* color mesh by own defined color
[-]label
* color labels separately
[.]by atom
* set label color by atom
[.]my color
* color labels by own defined color
[-]stick
* color sticks separately
[.]standard
* set stick color to standard
[.]my color
* color sticks by own defined color
[-]my colors
* use/append own defined colors
* own colors can be defined by
* Setting->Colors..->New
* to keep color settings for
* other pymol sessions
* you have to set the colors
* in .pymolrc or similar
* pymol setting file
* like
* set_color mycolor,[ 1.00, 1.00, 1.00]
[+]show
[-]surface flag
* set surface flag of atoms to show hide
* or ignore fro surface calculation
[-]transparency
* set different transparency types
* on selections or atoms