GROMACS Plugin
| Type | PyMOL Plugin |
|---|---|
| Download | pymol_plugin_dynamics.py |
| Author(s) | Laboratory of Biomolecular Systems Simulations |
| License | GPLv3 |
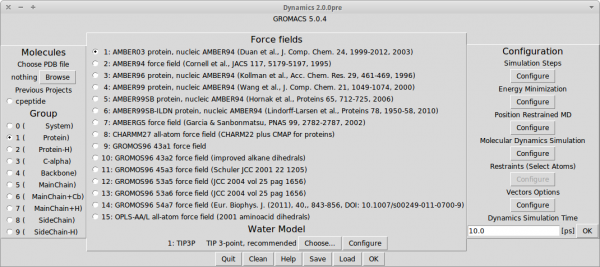
Dynamics PyMOL Plugin is plugin for PyMOL, which add molecular dynamics simulation feature. It is meant to be easy to use. The plugin uses GROMACS tools as a back-end. Project is developed as an open source and as such create full open source stack together with PyMOL and GROMACS. Software works on Linux, MacOS X and Windows/Cygwin.
Website
https://github.com/makson96/Dynamics
Features
- Easy to use GUI, to take advantage of complex software GROMACS.
- Work directly on molecules loaded to PyMOL.
- Display results of calculations directly in PyMOL.
- Set restraints, choose water models, force fields and many more.
- Save your work and finish calculations later or on the other machine.
- Do your work free of charge and without any restrictions. Feel free to modify any component of the stack.
- Minimum dependencies. Plugin use the same graphic libraries as PyMOL, so working PyMOL and GROMACS installations are enough to make plugin work.
Note that exact instruction of how to use program is in the software manual, which is available together with the plugin.
Installation
There are at least few ways to install the plugin. In this section we will describe the most common.
Ubuntu
The easiest way to install the plugin is to use PPA repositories:
sudo add-apt-repository ppa:makson96/dynamics sudo apt-get update sudo apt-get install pymol-plugin-dynamics
All required libraries will be downloaded as dependencies. Supported versions of Ubuntu are latest stable and latest LTS.
Gentoo
The plugin is also present in Gentoo ebuild system. To make the installation use (as root):
emerge --ask sci-chemistry/pymol-plugins-dynamics
Other Linux distributions and macOS
The second way for all other platforms, is to download latest release of the plugin directly from its GitHub webpage:
https://github.com/makson96/Dynamics/releases
Then run PyMOL as a root. On the top Menu choose Plugin->Manage Plugins->Install... Then choose pymol_plugin_dynamics.py file. After installation restart PyMOL with normal user privilege. On macOS you will need to use X11 Hybrid version of the PyMOL.
Required dependencies:
- PyMOL
- GROMACS
- ProDy (optional)
In order to take advantage of latest features you will need to have ProDy library installed.
Windows/Cygwin
- Download and install the latest version of Cygwin including appropriate code development packages.
- Download, compile, and install the latest version of GROMACS 2016.3 under Cygwin. Follow the standard compilation, installation and testing instructions to build, compile and install GROMACS 2016.3 under Cygwin. For example:
% cd gromacs-2016.3 % mkdir build % cmake .. -DBUILD_SHARED_LIBS=off -DGMX_BUILD_OWN_FFTW=ON -DCMAKE_INSTALL_PREFIX=/cygdrive/c/<YOUR_GROMACS_ROOT> % make >& make.log % make install >& make.install.log
- Test GROMACS from Cywin command line.
- Download and install 64 bit PyMOL 1.8.2.0, which comes with Python 2.7.9.
- Test PyMOL.
- Update PATH environment variable on Windows to include the following bin directories corresponding to GROMACS and Cygwin:
C:\<YOUR_GROMACS_ROOT>\bin C:\<YOUR_CYGWIN_PATH>\bin
- Install the GROMACS plugin following standard PyMOL instructions. Download latest plugin version from link: https://github.com/makson96/Dynamics/archive/master.zip and unpack it. Run PyMOL. On the top Menu choose Plugin->Manage Plugins->Install... Then choose pymol_plugin_dynamics.py file. After installation restart PyMOL
Latest Snapshots
You can download latest master snapshot by command:
git clone git://github.com/makson96/Dynamics.git
Then run PyMOL as a root. On the top Menu choose Plugin->Manage Plugins->Install... Then choose pymol_plugin_dynamics.py file. After installation restart PyMOL with normal user privilege.
Required dependencies:
- PyMOL
- GROMACS
- ProDy (optional)
- PLUMED (optional, compiled into GROMACS)
In order to take advantage of latest features you will need to have ProDy library installed and PLUMED compiled into GROMACS.
References
Molecular Dynamics Simulation by GROMACS Using GUI Plugin for PyMOL Tomasz Makarewicz and Rajmund Kaźmierkiewicz Journal of Chemical Information and Modeling 2013 53 (5), 1229-1234
Improvements in GROMACS plugin for PyMOL including implicit solvent simulations and displaying results of PCA analysis Tomasz Makarewicz, Rajmund Kaźmierkiewicz Journal of Molecular Modeling May 2016, 22:109
License
This software (including its Debian packaging) is available to you under the terms of the GPL-3, see "/usr/share/common-licenses/GPL-3". Software is created and maintained by Laboratory of Biomolecular Systems Simulation at University of Gdansk.
Contributors:
- Tomasz Makarewicz
- Ajit B. Datta
- Sara Boch Kminikowska
- Manish Sud